Blue Graph¶
Unifying Python framework for graph analytics and co-occurrence analysis.

About¶
BlueGraph is a Python framework that consolidates graph analytics capabilities from different graph processing backends. It provides the following set of interfaces:
preprocessing and co-occurrence analysis API providing semantic property encoders and co-occurrence graph generators;
graph analytics API providing interfaces for computing graph metrics, performing path search and community detection;
representation learning API for applying various graph embedding techniques;
representation learning downstream tasks API allowing the user to perform node classification, similarity queries, link prediction.
Using the built-in PGFrame data structure (currently, pandas-based implementation is available) for representing property graphs, it provides a backend-agnostic API supporting the following in-memory and persistent graph backends:
NetworkX (for the analytics API)
graph-tool (for the analytics API)
Neo4j (for the analytics and representation learning API);
StellarGraph (for the representation learning API).
gensim (for the representation learning API).
This repository originated from the Blue Brain effort on building a COVID-19-related knowledge graph from the CORD-19 dataset and analysing the generated graph to perform literature review of the role of glucose metabolism deregulations in the progression of COVID-19. For more details on how the knowledge graph is built, explored and analysed, see COVID-19 co-occurrence graph generation and analysis.
bluegraph package¶
BlueGraph’s API is built upon 4 main packages:
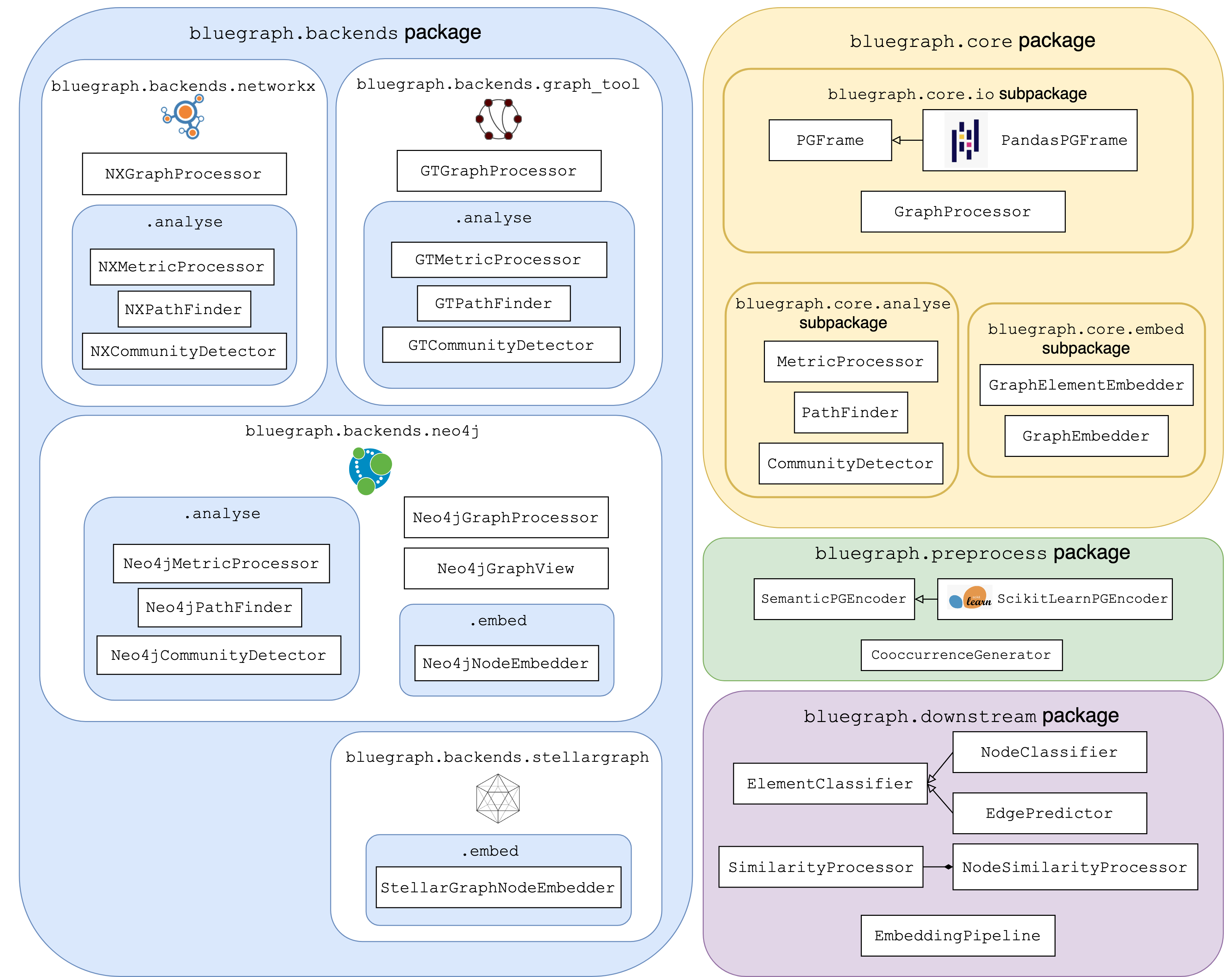
bluegraph.coreproviding the exchange data structure for graph representation that serves as the input to graph processors based on different backends (PGFrame), as well as basic interfaces for different graph analytics and embedding classes (MetricProcessor,PathFinder,CommunityDetector,GraphElementEmbedder, etc).bluegraph.backendsis a package that collects implementation of various graph processing and analytics interfaces for different graph backends (for example,NXPathFinderfor path search capabilities provided by NetworkX,Neo4jCommunityDetectorfor community detection methods provided by Neo4j, etc).bluegraph.preprocessis a package that contains utils for preprocessing property graphs (e.g.SemanticPGEncoderfor encoding node/edge properties as numerical vectors,CooccurrenceGeneratorfor generation and analysis of co-occurrence relations in PGFrames.)bluegraph.downstreamis a package that provides a set of utils for various downstream tasks based on vector representations of graphs and graph elements (for example,NodeSimilarityProcessorfor building and querying node similarity indices based on vector representation of nodes,EdgePredictorfor predicting true and false edges of the graph based on vector representation of its nodes,EmbeddingPipelinefor stacking pipelines of graph preprocessing, embedding, similarity index building, etc).
Main components of BlueGraph’s API are illustrated in the following diagram:
cord19kg package¶
The cord19kg package contains a set of tools for interactive exploration and analysis of the CORD-19 dataset using the co-occurrence analysis of the extracted named entities. It includes data preparation and curation helpers, tools for generation and analysis of co-occurrence graphs. Moreover, it provides several interactive mini-applications (based on JupyterDash and ipywidgets) for Jupyter notebooks allowing the user to interactively perform:
entity curation;
graph visualization and analysis;
dataset saving/loading from Nexus.
services package¶
Collects services included as a part of BlueGraph. Currently, only a mini-service for retrieving embedding vectors and similarity computation is included as a part of this repository (see embedder service specific README).
Installation¶
It is recommended to use a virtual environment such as venv or conda environment.
Installing backend dependencies¶
If you want to use graph-tool as a backend, you need to manually install the library (it cannot be simply installed by running pip install), as it is not an ordinary Python library, but a wrapper around a C++ library (please, see graph-tool installation instructions). Currently, BlueGraph supports graph-tool<=2.37.
Similarly, if you want to use the bluegraph.downstream.similarity module for building similarity indices (on embedded nodes, for example), you should install the Facebook Faiss library separately. Please, see Faiss installation instructions (conda and conda-forge installation available).
You can install both graph-tool and the Facebook Faiss library by creating a new environment with the right dependencies using conda, as follows:
conda create --name <your_environment> -c conda-forge graph-tool==2.37 faiss python=<your_python>
conda activate <your_environment>
The same holds for the Neo4j backend: in order to use it, the database should be installed and started (please, see Neo4j installation instructions). Typically, the Neo4j-based interfaces provided by BlueGraph require the database uri (the bolt port), username and password to be provided. In addition, BlueGraph uses the Neo4j Graph Data Science (GDS) library, which should be installed separately for the database on which you would like to run the analytics (see installation instructions). Current supported Neo4j GDS version is >=1.6.1.
Installing BlueGraph¶
BlueGraph supports Python versions >= 3.7 and pip >= 21.0.1. To update pip from the older versions run:
pip install --upgrade pip wheel setuptools
The stable version of BlueGraph can be installed from PyPI using:
pip install bluegraph
The development version of BlueGraph can be installed from the source by cloning the current repository as follows:
git clone https://github.com/BlueBrain/BlueGraph.git
cd BlueGraph
Basic version including only the NetworkX backend can be installed using:
pip install bluegraph
The prerequisites for using the graph-tool backend can be found in ‘Installing backend dependencies’. You can also install additional backends for Neo4j and StellarGraph by running the following:
pip install bluegraph[<backend>]
Where <backend> has one of the following values neo4j or stellargraph.
Alternatively, a version supporting all the backends can be installed by running the following commands:
pip install bluegraph[all]
In order to use the cord19kg package and its interactive Jupyter applications, run:
pip install bluegraph[cord19kg]
Getting started¶
The examples directory contains a set of Jupyter notebooks providing tutorials and usecases for BlueGraph.
To get started with property graph data structure PGFrame provided by BlueGraph, get an example of semantic property encoding, see the PGFrames and semantic encoding tutorial notebook.
To get familiar with the ideas behind the co-occurrence analysis and the graph analytics interface provided by BlueGraph we recommend to run the following example notebooks:
Literature exploration (PGFrames + in-memory analytics tutorial) illustrates how to use BlueGraphs’s analytics API for in-memory graph backends based on the
NetworkXand thegraph-toollibraries.NASA keywords (PGFrames + Neo4j analytics tutorial) illustrates how to use the Neo4j-based analytics API for persistent property graphs.
Embedding and downstream tasks tutorial starts from the co-occurrence graph generation example and guides the user through the graph representation learning and all it’s downstream tasks including node similarity queries, node classification and edge prediction.
Create and run embedding pipelines illustrates how embedding pipelines can be built and executed using BlueGraph.
Finally, Create and push embedding pipeline into Nexus.ipynb illustrates how embedding pipelines can be created and pushed to Nexus and Embedding service API shows how embedding service that retrieves the embedding pipelines from Nexus can be used.
Getting started with cord19kg¶
The cord19kg packages provides examples of CORD-19-specific co-occurrence analysis. Please, see more details on the CORD-19 analysis and exploration pipeline of the Blue Brain Project here.
We recommend starting from the Co-occurrence analysis tutorial notebook providing a simple starting example.
The Topic-centered co-occurrence network analysis of CORD-19 notebook provides a full analysis pipeline on the selection of 3000 articles obtained by searching the CORD-19 dataset using the query “Glucose is a risk factor for COVID-19” (the search is performed using BlueBrainSearch).
The Nexus-hosted co-occurrence network analysis of CORD-19 notebook provides an example for the previously mentioned 3000-article dataset, where datasets corresponding to different analysis steps can be saved and loaded to and from a Blue Brain Nexus project.
Finally, the generate_10000_network.py script allows the user to generate the co-occurrence networks for 10’000 most frequent entities extracted from the entire CORD-19v47 database (based on paper- and paragraph- level entity co-occurrence). To run the script, simply execute python generate_10000_network.py from the examples folder.
Note that the generated networks are highly dense (contain a large number of edges, for example, ~44M edges for the paper-based network), and the process of their generation, even if parallelized, is highly costly.
Licensing¶
Blue Graph is distributed under the Apache 2 license.
Included example scripts and notebooks (BlueGraph/examples and BlueGraph/cord19kg/examples) are distributed under the 3-Clause BSD License.
Data files stored in the repository are distributed under the Commons Attribution 4.0 International license (CC BY 4.0) License.
Funding & Acknowledgements¶
The development of this project was supported by funding to the Blue Brain Project, a research center of the École polytechnique fédérale de Lausanne (EPFL), from the Swiss government’s ETH Board of the Swiss Federal Institutes of Technology.
Copyright (c) 2020-2022 Blue Brain Project/EPFL